Appearance
Navigating Pantograph
Pantograph offers intuitive scrolling (by dragging) and zooming (by mouse wheel) within each loaded zoom level.
Zoom levels
The user can switch between zoom levels with the "Bin width" dropdown in the toolbar, or with right-clicking on any column and selecting "Zoom in" or "Zoom out". This will change to the next available zoom levels (e.g. from 1 to 10).
TIP
If a column is highlighted (i.e. right-click selected), changing to an arbitrary zoom level via the dropdown menu in the toolbar will center and highlight that position in the new zoom level as well. This way, the user can better follow the current position after a zoom level change.
Navigate to positions and genes
The Navigation Menu in the toolbar allows to quickly navigate to path-specific positions, to gene locations, or to a pangenome region.
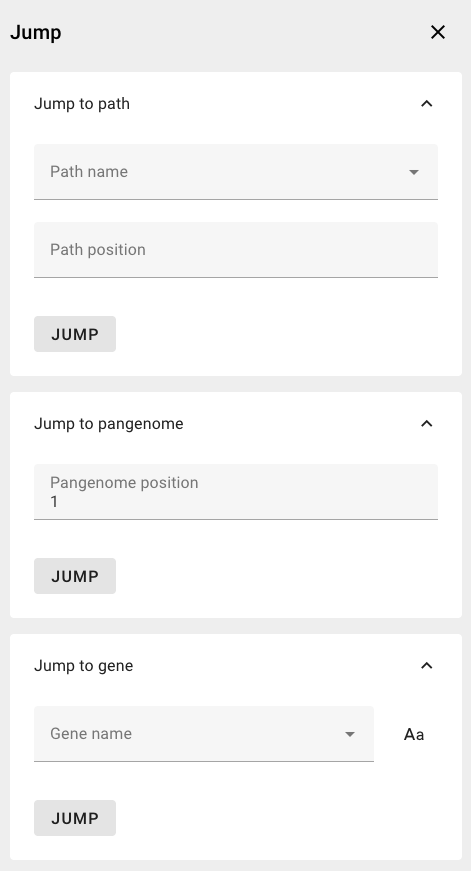
- Jump to path: Navigate to path-specific coordinates
Specifying a path and a position on that path will navigate to and highlight the position in the pangenome - Jump to pangenome: Navigate to and highlight a position in the pangenome sequence
- Jump to gene: Navigate to and highlight the start bin of a gene
Please note: The bin width is automatically set to 100, if the current bin width is larger than 100, for better identification of the target gene
Navigating Links
Links can be followed to their other end in lower zoom levels by right clicking on them and selecting "Follow link". The pangenome region of the other end is loaded and the bin is highlighted where the path continues in the tracks that follow this link.
User Interaction
Sorting Tracks
There are following possibilities to sort tracks:
- by genetic variation of the graph tracks or variant tracks in a single column:
Right click on the column to sort by and select "Sort graph paths" or "Sort variant tracks". - by any meta data (numeric or categorical; automatically detected):
Select the meta data category in the Toolbar's Sort Menu, or click on the little arrow to the top of the meta data column (shows up on mouse over). Repeated clicking will toggle between ascending/descending sort order. - by path name (alphabetically):
Select "Name" in the Toolbar's Sort Menu, or click on the little sorting arrow on top of the path names (shows up on mouse over).
Right-click Menu
Right-clicking on any column highlights it and opens the right click menu with the following options:
- Plot by metadata (graph paths): Plot phenotypic distributions stratified by genetic data
- Plot by metadata (variants) (if variant tracks are displayed): Plot phenotypic distributions stratified by genotypes
- Sort graph paths
The graph tracks get sorted by the coverage values in the highlighted column. - Sort variant tracks (if variant tracks are displayed)
The variant tracks get sorted by their genotyping values in the highlighted column. If more than one variant is present in a column, the tracks get sorted by the first variant. - Zoom in/out
The immediate next higher or lower zoom level, if available, is loaded, centering the view at the highlighted column. - Follow Link (only shown at link columns)
The view navigates to the other end of the link in the pangenome graph, highlighting the immediate next sequence column according to the traversal through the graph.
Tooltips
Tooltips over graph tracks denote the track name and the fraction of present bases in the given column (window of the pangenome sequence) of the path (between 0 and 1), shorthand: "Cov.". At inverted columns, the fraction of inverted bases from the whole window is additionally indicated ("Inv."). If all bases in the window are inverted, Cov and Inv values are equal. More info here.
Tooltips over genic regions additionally indicate the gene name (retrieved from the "ID" field of gff3 files), the strand, and the exon number.
Tooltips over variant tracks indicate the track name, the reference and position of a variant and genotype and allele information, if available. If more than 5 variants are within the bin, only the number of variants is displayed.